Application Note with bit.bio & Charles River Laboratories
This application note resulted from a collaboration between MaxWell Biosystems, bit.bio and Charles River Laboratories (CRL). In this study, CRL functionally characterised ioGlutamatergic Neurons HTT 50CAG/WT and ioGlutamatergic Neurons Huntington’s disease model developed by bit.bio using the MaxTwo HD-MEA system.
With this application note, you can learn how using the MaxTwo HD-MEA system and MaxLab Live Software Assays, such as the AxonTracking Assay, CRL generated phenotypic data showing that bit.bio’s Huntington’s disease model has delayed neuronal network formation, decreased axonal branching and decreased spontaneous activity compared to the isogenic control.
Title
Developing next-generation in vitro phenotypic assays for Huntington’s disease by combining a precision reprogrammed hiPSC-derived disease model with high-density microelectrode arrays
Abstract
Huntington’s disease (HD) is a fatal neurodegenerative disorder caused by the expansion of the CAG trinucleotide repeat in the huntingtin (HTT) gene leading to the expression of a mutant protein (mHTT), which affects neuronal health and function. There is currently no effective cure or treatment to slow down or stop the disease’s progression. One reason for this is the lack of accurate and easy-to- use HD models that recapitulate the phenotypes and pathology seen in patients. Additionally, more advanced methods are required to record and analyze functional properties of cultured neurons over the entire maturation period, in order to gain insights into the underlying mechanisms of neurodegenerative diseases. In this Application Note, Charles River Laboratories (Cambridge, UK) functionally characterisedioGlutamatergic Neurons HTT 50CAG/WT and ioGlutamatergic Neurons developed by bit.bio (Cambridge, UK). Together both cell types form a genetically matched isogenic disease model for the modelling of HD in vitro. For this purpose, they used the MaxTwo high-density microelectrode array platform developed by MaxWell Biosystems (Zürich, Switzerland). This work shows several HD-relevant electrophysiological differences and altered axonal function in the disease model cells when compared to the wild-type control.
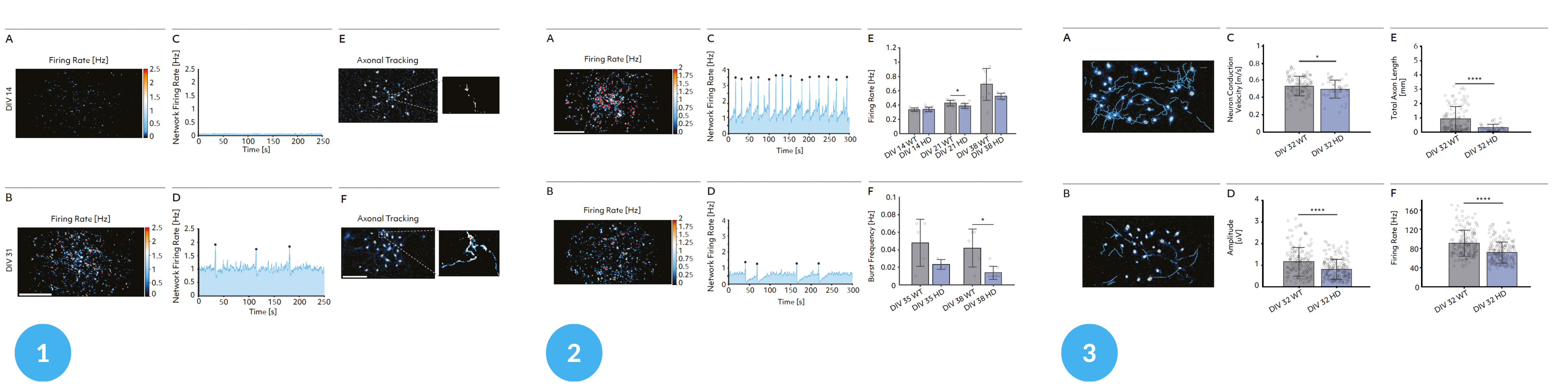

1 – Maturation analysis; 2 – Single-cell and network development analysis; 3 – Single-cell analysis
 English
English


