Getting Started with Closed-Loop Experiments on HD-MEAs
Author
Date
Tags
What is biocomputing?
Unlike traditional processors that rely on fixed circuitry and binary bits, biocomputing uses living networks, neuronal cultures or organoids, to process information through activity-dependent adaptation. Think of it as an artificial intelligence (AI) system not only inspired by neural networks but one that can draw insights directly from living networks, one that learn continuously and handles ambiguity with nuance – blending the gap between artificial and natural intelligence.
In recent years, there has been media coverage from the BBC, New York Times, and Bloomberg of a resurgence in biocomputing – sparked by our High-Density Microelectrode Array (HD-MEA) technology and leveraged by our users to force in vitro models of the brain to perform computational tasks. Across the world, our user community trained cultured neurons to improve their performance in a game of Pong (Kagan et al., 2022), trained organoids to balance a cartpole (Robbins et al., 2026) as well as recognize different speech patterns (Cai et al., 2023), and interfaced organoids with artificial networks to emulate disease models as well as guide therapeutic interventions (Beaubois et al., 2024).
In this blog post, we’ll take one step back to the building blocks: how to implement a simple closed-loop stimulation paradigm using the MaxLab Live API, so you can go from “record and stimulate” to a working feedback loop that can be extended toward training paradigms and embodied tasks.
How closed-loop feedback can induce learning
Neurons – the building blocks of the brain – are plastic and have an extraordinary capacity to adapt. Patterns of electrical activity can drive adaptation across multiple scales, leading to adjustments in synaptic strength, intrinsic excitability, and overall network dynamics. Over time, these activity-dependent adjustments enable learning through functional, and sometimes even structural changes.
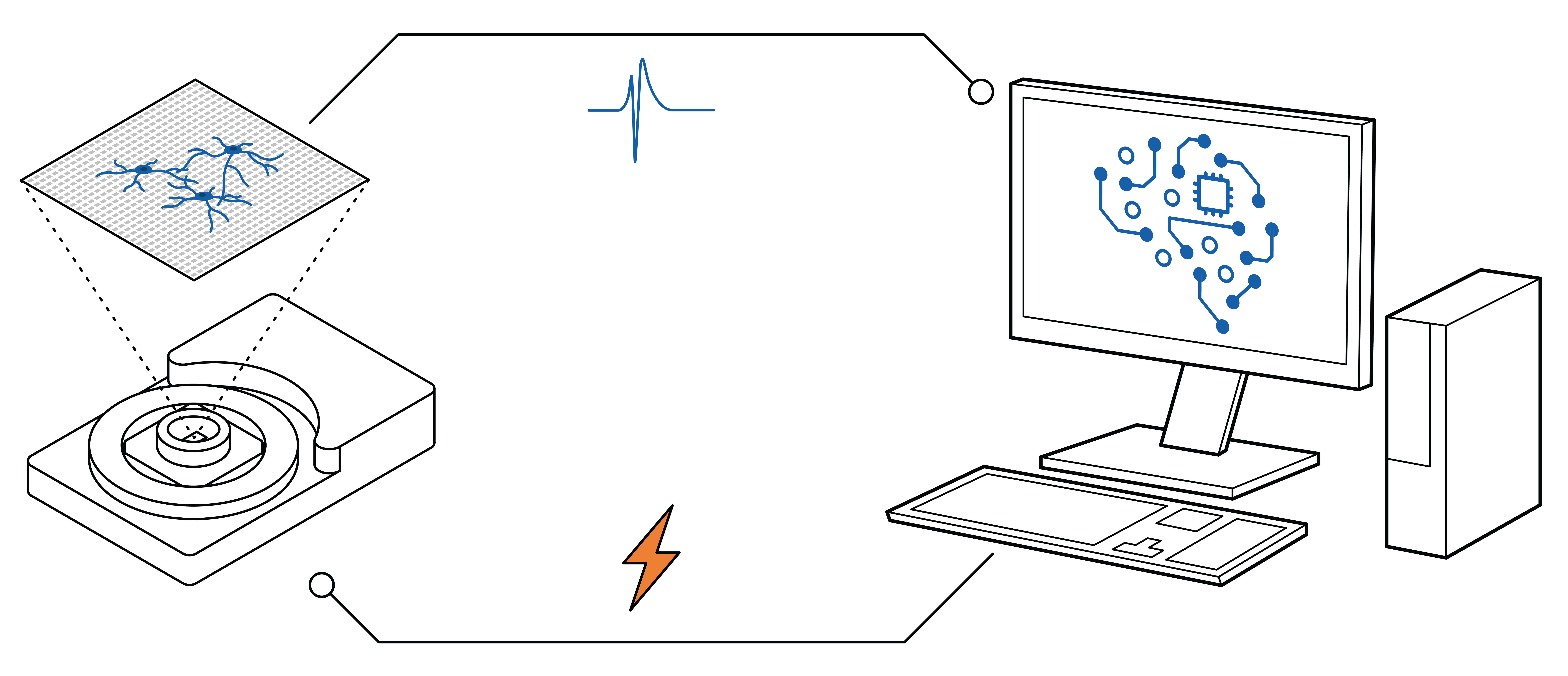
MaxWell Biosystems High-Density Microelectrode Arrays (HD-MEAs) make it possible to observe and interact with biological neuronal networks (BNNs) in real time. This enables low-latency closed-loop paradigms where neural activity is continuously measured, interpreted, and shaped through feedback. While implementations vary widely, most closed-loop systems follow the same steps:
- Input encoding: Represent the current state of the task or environment in a stimulus pattern delivered to the BNN.
- Observe: Record spiking activity from the BNN using a set of electrodes.
- Evaluate: Decode the activity into task-relevant variables (or a performance score) and determine the next intervention.
- Feedback: Deliver a distinct stimulation signal to shape future responses, for example by reinforcing desired behaviour or signalling error.
In other words, closed-loop paradigms often involve two different ways of stimulating the network. One stimulus encodes information about the current task or environment and serves as input to the BNN. The resulting neural activity is then recorded and decoded into an action or performance metric. A second, distinct feedback signal can then be applied to reinforce desired behaviour or otherwise guide the network’s adaptation over time.
A reproducible closed-loop example
To illustrate the basic principles of closed-loop control with the MaxLab Live API, we use a simple example in which the relative timing of two input events determines which output is stimulated. Spike-like events are generated on two input channels, A and B, and their temporal difference, Δt(A,B), is evaluated in real time. Depending on this delay, the system triggers stimulation on either output C or output D. This creates a minimal and intuitive demonstration of how activity can be observed, interpreted, and used to drive feedback.
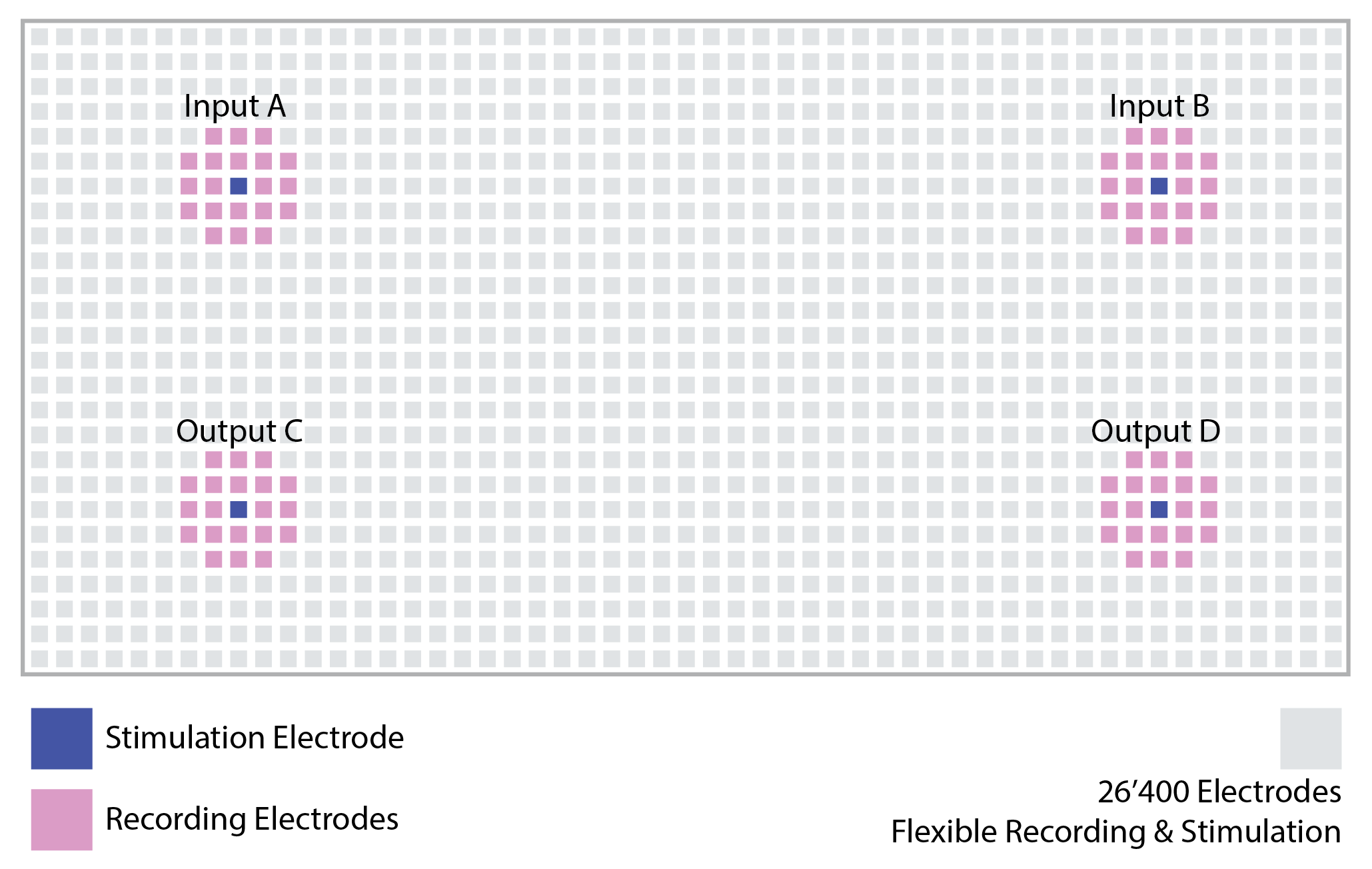
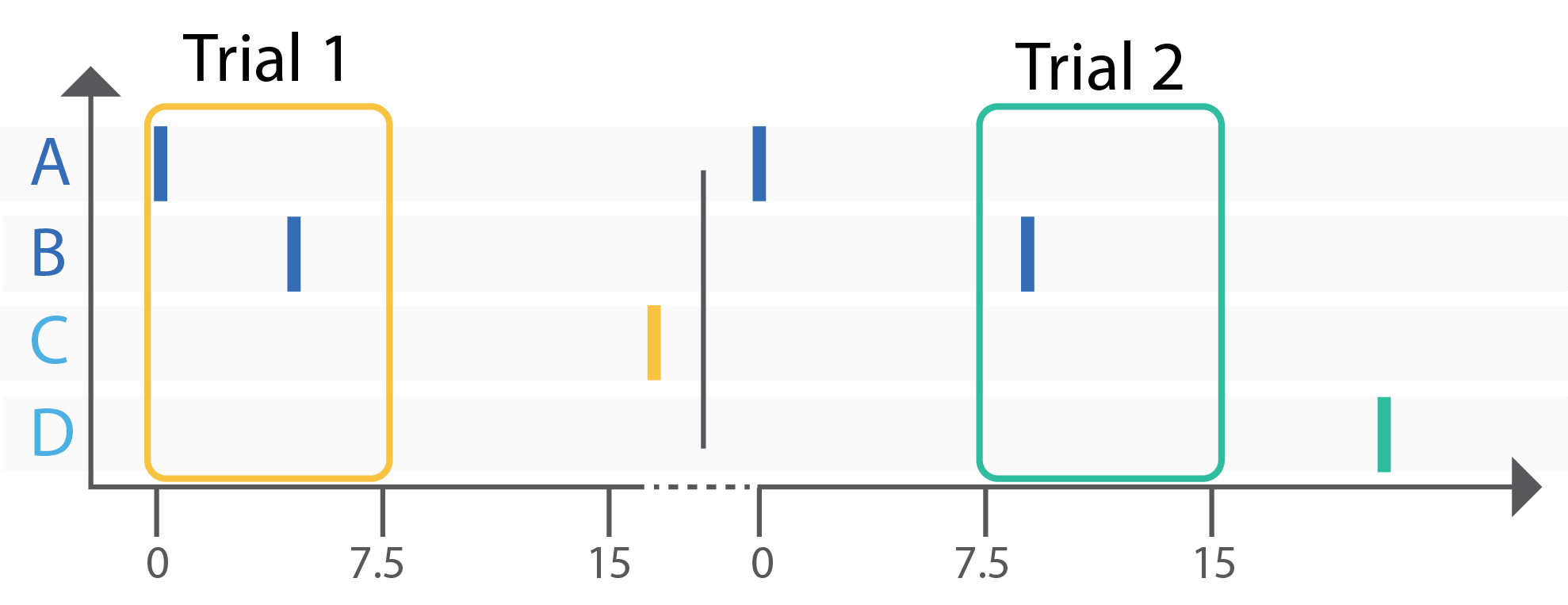
To make this tutorial easy to replicate and understand, we intentionally run it on a chip filled with saline buffer rather than a neuronal culture. In saline, there are no biological spikes, so we generate spike-like events by stimulating electrodes near our input electrodes. These events are easy to produce and repeat, easy to visualize in real time, and make the resulting input-output behaviour immediately intuitive. This allows you to validate the complete closed-loop pipeline before moving on to real biological activity.
To reproduce this example, you need a MaxOne Single-Well HD-MEA system connected to a computer running MaxLab Live, together with a MaxOne HD-MEA chip filled with saline buffer. The implementation is split into two parts: a Python module that sets up the experiment, defines electrodes and stimulation patterns, and runs the recording; and a C++ module that monitors activity with low latency and triggers stimulation according to a simple rule.
Step 1 – Configure the experiment in Python
First, use Python to set up the experiment. This includes selecting and preparing the recording and stimulation electrodes, defining the stimulation patterns, and starting the recording.
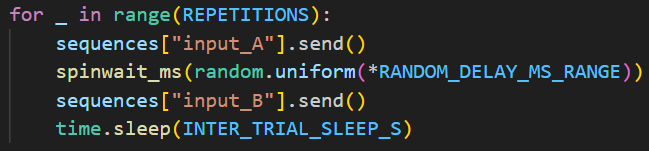
Step 2 - Generate the “input events” in Python
Next, we create controllable inputs by stimulating electrodes near A and B in a timed loop, producing spike-like events on A and B with varying inter-event delays and therefore a controlled range of Δt(A,B) values over time.
Step 3 - Implement the real-time loop in C++
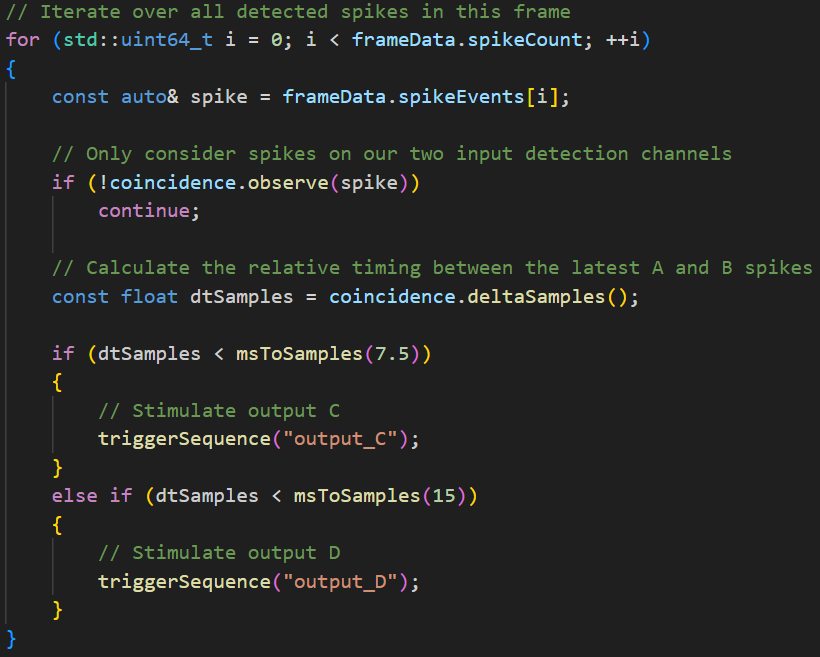
The C++ module implements the low-latency closed-loop logic. It monitors activity on the input electrodes A and B in real time, tracks the time difference between their events, and applies the stimulation rule:
- If Δt(A,B) ≤ 7.5 ms, stimulate electrode C.
- If 7.5 ms < Δt(A,B) , stimulate electrode D.
Step 4 - Visualize while it runs
While both scripts are running, use MaxLab Live Scope to inspect the signals in real time and confirm:
- Spike-like events on A and B (stimulus-evoked artifacts), and
- Conditional stimulation on C or D depending on Δt(A,B).
Generalizing to a real-world example
The example above is intentionally simple, but the same framework extends to more complex paradigms. For example, Robbins and colleagues (2026) at the University of California, Santa Cruz, built a closed-loop paradigm to train organoids to balance a cartpole. In that work, they developed an analysis pipeline to identify candidate neural units and connectivity, then selected encoding/decoding/training channels. They used rate coding for input (encoding) and output (decoding) and showed that adaptive training outperformed random or null paradigms. For more information on this fascinating work, check out our recent blog post.
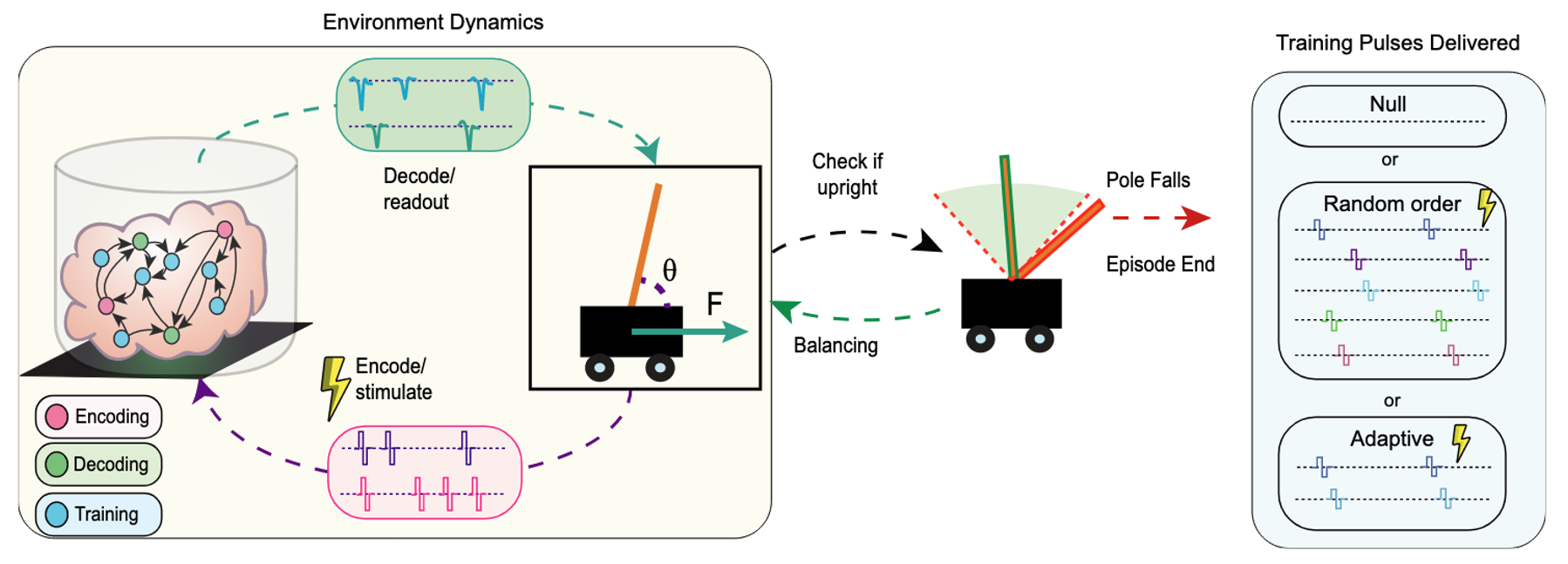
CC-BY-NC 4.0.
Summary and outlook
This blog is part of a series on neurocomputing. Here, we demonstrated a simple closed-loop paradigm and how it can be implemented on an HD-MEA system using the MaxLab Live API.
At the same time, the broader topic of “training” living networks is controversial: ethical implications and the risk of inflated claims are actively debated across scientific, governmental, ethical, legal, and business communities. (In this series, we’ll aim to be explicit about what the data show, what they do not show, and where interpretations differ.)
In upcoming posts, we’ll highlight voices from the user community—both pioneers and researchers pushing the field forward—and we’ll build on this example to illustrate richer paradigms and use cases.
Join our biocomputing user community—maybe you’ll become part of turning what once felt like science fiction into practical, reproducible experiments.
Get the code
Want to reproduce this closed-loop demo on your own setup?
We’re happy to share the Python and C++ example code, along with notes on electrode configuration and recommended Scope views. Reach out to us via our Contact us page (or your local MaxWell representative) and mention “Closed-loop demo code” in the message.
Related
Resources
MaxOne
Versatility and functionality in one compact device

MaxLab Live
All-in-one Software
Biocomputing

Neuronal Cell Cultures
Organoids
Goal-directed learning in cortical organoids

In vitro neurons learn and exhibit sentience when embodied in a simulated game-world

Brain organoid reservoir computing for artificial intelligence

BiœmuS: A new tool for neurological disorders studies through real-time emulation and hybridization using biomimetic Spiking Neural Network


%2520Li.avif)