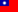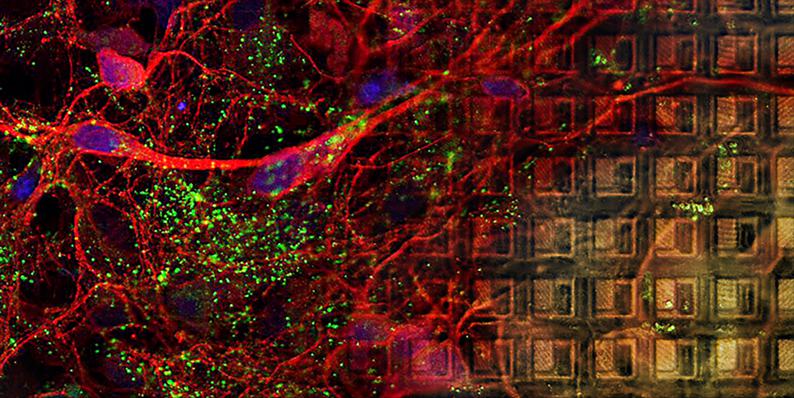

Primary Cell Cultures on MaxOne MEA
Dissociated cell cultures can be plated, grown, and imaged on MaxOne MEAs. The figure shows an overlay of multiple confocal stack images: cell bodies and neurites are visible on the left side and the electrode array is focused on the right side.
What You Can Do
Single Cell Level Analysis
Isolate extracellular action potentials from individual cells easily.
Select electrodes with the best signals from each cell and improve your spike sorting.


Analyze subcellular features, such as full axonal arbors in single neurons.
Electrical signals tracked along neurites enable to investigate novel parameters, difficult to obtain with previous technologies.


High-resolution axonal action potential propagation tracking
The video shows the propagation of an AP through the axonal arbor of a single neuron. Electrical signals tracked along neurites enable to investigate novel parameters, difficult to obtain with previous technologies.
Axonal branch signals tracking
The video shows the propagation of a stimulation-induced action potential through a branching segment of an axon. To reduce noise and better reveal the action potential, 300 trials have been averaged.
Network Level Analysis
Record and/or stimulate every cell on the MEA.
MaxOne allows simultaneous recording from thousands of cells yields rich datasets for studying neuronal networks. Stimulation experiments can be designed with high flexibility to manipulate network dynamics.
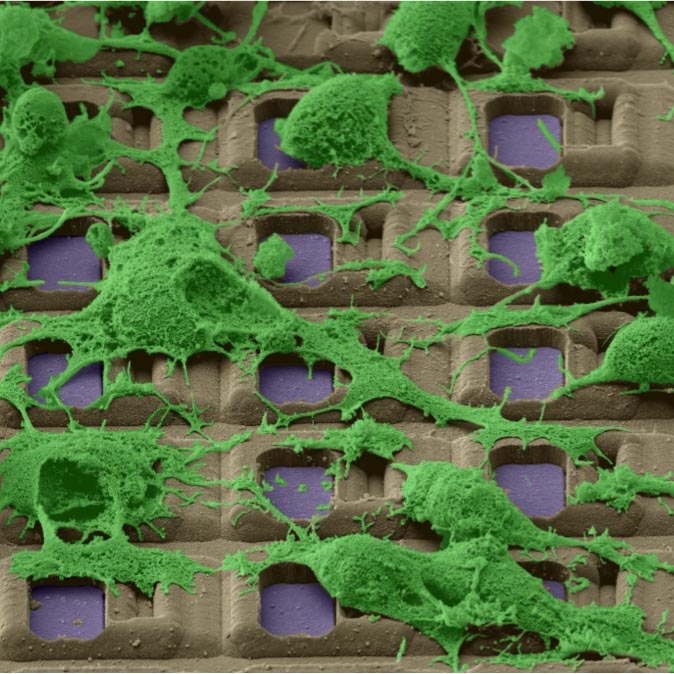

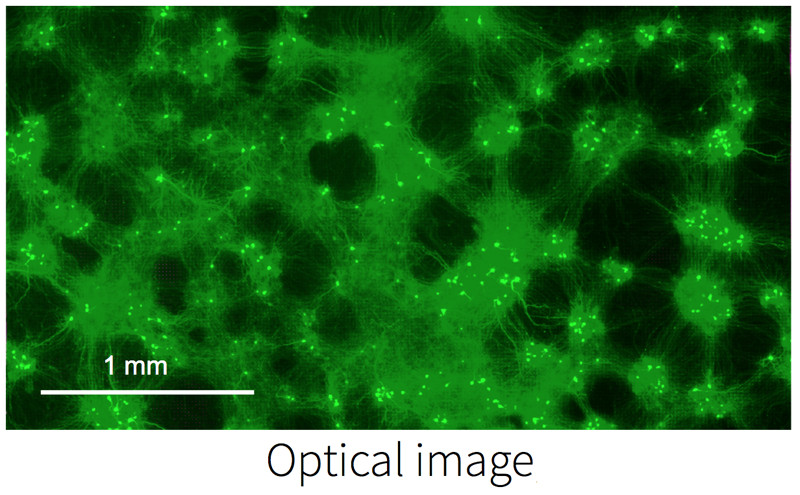
Electrically image active cells on the MEA.
Localize active cells on the MEA during experiments. The optical image from fluorescence microscopy matches the electrical image obtained using MaxOne.




Study cells and networks across different spatial and temporal scales.
Action potentials and local field potentials can be simultaneously recorded by MaxOne over time scales of microseconds to months. The raster plot shows the dynamics of network activity using 1,024 active electrode sites.


Selected Publications
  | Hornauer, Philipp; Prack, Gustavo; Anastasi, Nadia; Ronchi, Silvia; Kim, Taehoon; Donner, Christian; Fiscella, Michele; Borgwardt, Karsten; Taylor, Verdon; Jagasia, Ravi; Roqueiro, Damian; Hierlemann, Andreas; Schröter, Manuel DeePhys: A machine learning–assisted platform for electrophysiological phenotyping of human neuronal networks Journal Article Stem Cell Reports, 2024. @article{Hornauer2024, title = {DeePhys: A machine learning–assisted platform for electrophysiological phenotyping of human neuronal networks}, author = {Philipp Hornauer and Gustavo Prack and Nadia Anastasi and Silvia Ronchi and Taehoon Kim and Christian Donner and Michele Fiscella and Karsten Borgwardt and Verdon Taylor and Ravi Jagasia and Damian Roqueiro and Andreas Hierlemann and Manuel Schröter}, url = {https://www.sciencedirect.com/science/article/pii/S2213671123005015}, doi = {10.1016/j.stemcr.2023.12.008}, year = {2024}, date = {2024-01-25}, journal = {Stem Cell Reports}, abstract = {Reproducible functional assays to study in vitro neuronal networks represent an important cornerstone in the quest to develop physiologically relevant cellular models of human diseases. Here, we introduce DeePhys, a MATLAB-based analysis tool for data-driven functional phenotyping of in vitro neuronal cultures recorded by high-density microelectrode arrays. DeePhys is a modular workflow that offers a range of techniques to extract features from spike-sorted data, allowing for the examination of functional phenotypes both at the individual cell and network levels, as well as across development. In addition, DeePhys incorporates the capability to integrate novel features and to use machine-learning-assisted approaches, which facilitates a comprehensive evaluation of pharmacological interventions. To illustrate its practical application, we apply DeePhys to human induced pluripotent stem cell–derived dopaminergic neurons obtained from both patients and healthy individuals and showcase how DeePhys enables phenotypic screenings.}, keywords = {}, pubstate = {published}, tppubtype = {article} } Reproducible functional assays to study in vitro neuronal networks represent an important cornerstone in the quest to develop physiologically relevant cellular models of human diseases. Here, we introduce DeePhys, a MATLAB-based analysis tool for data-driven functional phenotyping of in vitro neuronal cultures recorded by high-density microelectrode arrays. DeePhys is a modular workflow that offers a range of techniques to extract features from spike-sorted data, allowing for the examination of functional phenotypes both at the individual cell and network levels, as well as across development. In addition, DeePhys incorporates the capability to integrate novel features and to use machine-learning-assisted approaches, which facilitates a comprehensive evaluation of pharmacological interventions. To illustrate its practical application, we apply DeePhys to human induced pluripotent stem cell–derived dopaminergic neurons obtained from both patients and healthy individuals and showcase how DeePhys enables phenotypic screenings. |
  | Silverman, Jill L; Fenton, Timothy; Haouchine, Olivia; Hallam, Elizabeth; Smith, Emily; Jackson, Kiya; Rahbarian, Darlene; Canales, Cesar; Adhikari, Anna; Nord, Alex; Ben-Shalom, Roy Hyperexcitability and translational phenotypes in a preclinical model of SYNGAP1mutations Journal Article Research Square, 2023. @article{Silverman2023, title = {Hyperexcitability and translational phenotypes in a preclinical model of SYNGAP1mutations}, author = {Jill L. Silverman and Timothy Fenton and Olivia Haouchine and Elizabeth Hallam and Emily Smith and Kiya Jackson and Darlene Rahbarian and Cesar Canales and Anna Adhikari and Alex Nord and Roy Ben-Shalom}, url = {https://www.researchsquare.com/article/rs-3246655/v1}, doi = {https://doi.org/10.21203/rs.3.rs-3246655/v1}, year = {2023}, date = {2023-09-13}, journal = {Research Square}, abstract = {SYNGAP1is a critical gene for neuronal development, synaptic structure, and function. Although rare, the disruption of SYNGAP1directly causes a genetically identi able neurodevelopmental disorder (NDD) called SYNGAP1-related intellectual disability. Without functional SynGAP1 protein, patients present with intellectual disability, motor impairments, and epilepsy. Previous work using mouse models with a variety of germline and conditional mutations has helped delineate SynGAP1’s critical roles in neuronal structure and function, as well as key biochemical signaling pathways essential to synapse integrity. Homozygous loss of SYNGAP1is embryonically lethal. Heterozygous mutations of SynGAP1result in a broad range of phenotypes including increased locomotor activity, impaired working spatial memory, impaired cued fear memory, and increased stereotypic behavior. Ourinvivofunctional data, using the original germline mutation mouse line from the Huganir laboratory, corroborated robust hyperactivity and learning and memory de cits. Here, we describe impairments in the translational biomarker domain of sleep, characterized using neurophysiological data collected with wireless telemetric electroencephalography (EEG). We discoveredSyngap1+/− mice exhibited elevated spike trains in both number and duration, in addition to elevated power, most notably in the delta power band. Primary neurons fromSyngap1+/− mice displayed increased network ring activity, greater spikes per burst, and shorter inter-burst intervals between peaks using high density micro-electrode arrays (HD-MEA). This work is translational, innovative, and highly signi cant as it outlines functional impairments in Syngap1mutant mice. Simultaneously, the work utilized untethered, wireless neurophysiology that can discover potential biomarkers of Syngap1RID, for clinical trials, as it has done with other NDDs. Our work is substantial forward progress toward translational work for SynGAP1R-ID as it bridges in-vitroelectrophysiological neuronal activity and function with invivoneurophysiological brain activity and function. These data elucidate multiple quantitative, translational biomarkers invivoand invitrofor the development of treatments for SYNGAP1-related intellectual disability.}, keywords = {}, pubstate = {published}, tppubtype = {article} } SYNGAP1is a critical gene for neuronal development, synaptic structure, and function. Although rare, the disruption of SYNGAP1directly causes a genetically identi able neurodevelopmental disorder (NDD) called SYNGAP1-related intellectual disability. Without functional SynGAP1 protein, patients present with intellectual disability, motor impairments, and epilepsy. Previous work using mouse models with a variety of germline and conditional mutations has helped delineate SynGAP1’s critical roles in neuronal structure and function, as well as key biochemical signaling pathways essential to synapse integrity. Homozygous loss of SYNGAP1is embryonically lethal. Heterozygous mutations of SynGAP1result in a broad range of phenotypes including increased locomotor activity, impaired working spatial memory, impaired cued fear memory, and increased stereotypic behavior. Ourinvivofunctional data, using the original germline mutation mouse line from the Huganir laboratory, corroborated robust hyperactivity and learning and memory de cits. Here, we describe impairments in the translational biomarker domain of sleep, characterized using neurophysiological data collected with wireless telemetric electroencephalography (EEG). We discoveredSyngap1+/− mice exhibited elevated spike trains in both number and duration, in addition to elevated power, most notably in the delta power band. Primary neurons fromSyngap1+/− mice displayed increased network ring activity, greater spikes per burst, and shorter inter-burst intervals between peaks using high density micro-electrode arrays (HD-MEA). This work is translational, innovative, and highly signi cant as it outlines functional impairments in Syngap1mutant mice. Simultaneously, the work utilized untethered, wireless neurophysiology that can discover potential biomarkers of Syngap1RID, for clinical trials, as it has done with other NDDs. Our work is substantial forward progress toward translational work for SynGAP1R-ID as it bridges in-vitroelectrophysiological neuronal activity and function with invivoneurophysiological brain activity and function. These data elucidate multiple quantitative, translational biomarkers invivoand invitrofor the development of treatments for SYNGAP1-related intellectual disability. |
  | Habibollahi, Forough; Kagan, Brett J; Burkitt, Anthony N; French, Chris Critical dynamics arise during structured information presentation within embodied in vitro neuronal networks Journal Article Nature Communications, 2023. @article{Habibollahi2023, title = {Critical dynamics arise during structured information presentation within embodied in vitro neuronal networks}, author = {Forough Habibollahi and Brett J. Kagan and Anthony N. Burkitt and Chris French }, url = {https://www.nature.com/articles/s41467-023-41020-3}, doi = {https://doi.org/10.1038/s41467-023-41020-3}, year = {2023}, date = {2023-08-30}, journal = {Nature Communications}, abstract = {Understanding how brains process information is an incredibly difficult task. Amongst the metrics characterising information processing in the brain, observations of dynamic near-critical states have generated significant interest. However, theoretical and experimental limitations associated with human and animal models have precluded a definite answer about when and why neural criticality arises with links from attention, to cognition, and even to consciousness. To explore this topic, we used an in vitro neural network of cortical neurons that was trained to play a simplified game of ‘Pong’ to demonstrate Synthetic Biological Intelligence (SBI). We demonstrate that critical dynamics emerge when neural networks receive task-related structured sensory input, reorganizing the system to a near-critical state. Additionally, better task performance correlated with proximity to critical dynamics. However, criticality alone is insufficient for a neuronal network to demonstrate learning in the absence of additional information regarding the consequences of previous actions. These findings offer compelling support that neural criticality arises as a base feature of incoming structured information processing without the need for higher order cognition.}, keywords = {}, pubstate = {published}, tppubtype = {article} } Understanding how brains process information is an incredibly difficult task. Amongst the metrics characterising information processing in the brain, observations of dynamic near-critical states have generated significant interest. However, theoretical and experimental limitations associated with human and animal models have precluded a definite answer about when and why neural criticality arises with links from attention, to cognition, and even to consciousness. To explore this topic, we used an in vitro neural network of cortical neurons that was trained to play a simplified game of ‘Pong’ to demonstrate Synthetic Biological Intelligence (SBI). We demonstrate that critical dynamics emerge when neural networks receive task-related structured sensory input, reorganizing the system to a near-critical state. Additionally, better task performance correlated with proximity to critical dynamics. However, criticality alone is insufficient for a neuronal network to demonstrate learning in the absence of additional information regarding the consequences of previous actions. These findings offer compelling support that neural criticality arises as a base feature of incoming structured information processing without the need for higher order cognition. |
  | Radivojevic, Milos; Punga, Anna Rostedt Functional imaging of conduction dynamics in cortical and spinal axons Journal Article eLife, 2023. @article{Radivojevic2023_2, title = {Functional imaging of conduction dynamics in cortical and spinal axons}, author = {Milos Radivojevic and Anna Rostedt Punga}, url = {https://elifesciences.org/articles/86512}, doi = {https://doi.org/10.7554/eLife.86512}, year = {2023}, date = {2023-08-22}, journal = {eLife}, abstract = {Mammalian axons are specialized for transmitting action potentials to targets within the central and peripheral nervous system. A growing body of evidence suggests that, besides signal conduction, axons play essential roles in neural information processing, and their malfunctions are common hallmarks of neurodegenerative diseases. The technologies available to study axonal function and structure integrally limit the comprehension of axon neurobiology. High-density microelectrode arrays (HD-MEAs) allow for accessing axonal action potentials at high spatiotemporal resolution, but provide no insights on axonal morphology. Here, we demonstrate a method for electrical visualization of axonal morphologies based on extracellular action potentials recorded from cortical and motor neurons using HD-MEAs. The method enabled us to reconstruct up to 5-cm-long axonal arbors and directly monitor axonal conduction across thousands of recording sites. We reconstructed 1.86 m of cortical and spinal axons in total and found specific features in their structure and function.}, keywords = {}, pubstate = {published}, tppubtype = {article} } Mammalian axons are specialized for transmitting action potentials to targets within the central and peripheral nervous system. A growing body of evidence suggests that, besides signal conduction, axons play essential roles in neural information processing, and their malfunctions are common hallmarks of neurodegenerative diseases. The technologies available to study axonal function and structure integrally limit the comprehension of axon neurobiology. High-density microelectrode arrays (HD-MEAs) allow for accessing axonal action potentials at high spatiotemporal resolution, but provide no insights on axonal morphology. Here, we demonstrate a method for electrical visualization of axonal morphologies based on extracellular action potentials recorded from cortical and motor neurons using HD-MEAs. The method enabled us to reconstruct up to 5-cm-long axonal arbors and directly monitor axonal conduction across thousands of recording sites. We reconstructed 1.86 m of cortical and spinal axons in total and found specific features in their structure and function. |
  | Duru, Jens; Maurer, Benedikt; Doran, Ciara Giles; Jelitto, Robert; Küchler, Joël; Ihle, Stephan J; Ruff, Tobias; John, Robert; Genocchi, Barbara; Vörös, János Investigation of the input-output relationship of engineered neural networks using high-density microelectrode arrays Journal Article Biosensors and Bioelectronics, 2023. @article{Duru2023b, title = {Investigation of the input-output relationship of engineered neural networks using high-density microelectrode arrays}, author = {Jens Duru and Benedikt Maurer and Ciara Giles Doran and Robert Jelitto and Joël Küchler and Stephan J. Ihle and Tobias Ruff and Robert John and Barbara Genocchi and János Vörös }, url = {https://www.sciencedirect.com/science/article/pii/S095656632300533X?via%3Dihub}, doi = {https://doi.org/10.1016/j.bios.2023.115591}, year = {2023}, date = {2023-08-18}, journal = {Biosensors and Bioelectronics}, abstract = {Bottom-up neuroscience utilizes small, engineered biological neural networks to study neuronal activity in systems of reduced complexity. We present a platform that establishes up to six independent networks formed by primary rat neurons on planar complementary metal–oxide–semiconductor (CMOS) microelectrode arrays (MEAs). We introduce an approach that allows repetitive stimulation and recording of network activity at any of the over 700 electrodes underlying a network. We demonstrate that the continuous application of a repetitive super-threshold stimulus yields a reproducible network answer within a 15 ms post-stimulus window. This response can be tracked with high spatiotemporal resolution across the whole extent of the network. Moreover, we show that the location of the stimulation plays a significant role in the networks' early response to the stimulus. By applying a stimulation pattern to all network-underlying electrodes in sequence, the sensitivity of the whole network to the stimulus can be visualized. We demonstrate that microchannels reduce the voltage stimulation threshold and induce the strongest network response. By varying the stimulation amplitude and frequency we reveal discrete network transition points. Finally, we introduce vector fields to follow stimulation-induced spike propagation pathways within the network. Overall we show that our defined neural networks on CMOS MEAs enable us to elicit highly reproducible activity patterns that can be precisely modulated by stimulation amplitude, stimulation frequency and the site of stimulation.}, keywords = {}, pubstate = {published}, tppubtype = {article} } Bottom-up neuroscience utilizes small, engineered biological neural networks to study neuronal activity in systems of reduced complexity. We present a platform that establishes up to six independent networks formed by primary rat neurons on planar complementary metal–oxide–semiconductor (CMOS) microelectrode arrays (MEAs). We introduce an approach that allows repetitive stimulation and recording of network activity at any of the over 700 electrodes underlying a network. We demonstrate that the continuous application of a repetitive super-threshold stimulus yields a reproducible network answer within a 15 ms post-stimulus window. This response can be tracked with high spatiotemporal resolution across the whole extent of the network. Moreover, we show that the location of the stimulation plays a significant role in the networks' early response to the stimulus. By applying a stimulation pattern to all network-underlying electrodes in sequence, the sensitivity of the whole network to the stimulus can be visualized. We demonstrate that microchannels reduce the voltage stimulation threshold and induce the strongest network response. By varying the stimulation amplitude and frequency we reveal discrete network transition points. Finally, we introduce vector fields to follow stimulation-induced spike propagation pathways within the network. Overall we show that our defined neural networks on CMOS MEAs enable us to elicit highly reproducible activity patterns that can be precisely modulated by stimulation amplitude, stimulation frequency and the site of stimulation. |
  | Yamamoto, Hideaki; Hirano-Iwata, Ayumi; Sato, Shigeo Microfluidic technologies for reconstituting neuronal network functions in vitro Journal Article JSAP Review, 2023. @article{Yamamoto2023, title = {Microfluidic technologies for reconstituting neuronal network functions in vitro}, author = {Hideaki Yamamoto and Ayumi Hirano-Iwata and Shigeo Sato}, url = {https://www.jstage.jst.go.jp/article/oubutsu/92/5/92_278/_article/-char/en}, doi = {10.11470/oubutsu.92.5_278}, year = {2023}, date = {2023-07-06}, journal = {JSAP Review}, abstract = {The structure and function of complex neuronal networks in the brain can be partially reconstituted in vitro by integrating cell culture and microfluidic device technologies. In this report, we review our recent studies on developing microfluidic devices to reconstitute small neuronal networks bearing a modular structure, which is a canonical structure found in the nervous systems of animals. We also describe the process of recording functional activity from the reconstituted neuronal networks. These fundamental technologies offer novel tools for investigating structure–function relationships in living neuronal networks and exploring the physical basis of biological computing in the brain.}, keywords = {}, pubstate = {published}, tppubtype = {article} } The structure and function of complex neuronal networks in the brain can be partially reconstituted in vitro by integrating cell culture and microfluidic device technologies. In this report, we review our recent studies on developing microfluidic devices to reconstitute small neuronal networks bearing a modular structure, which is a canonical structure found in the nervous systems of animals. We also describe the process of recording functional activity from the reconstituted neuronal networks. These fundamental technologies offer novel tools for investigating structure–function relationships in living neuronal networks and exploring the physical basis of biological computing in the brain. |
  | Girardi, Gregory; Zumpano, Danielle; Goshi, Noah; Raybould, Helen; Seker, Erkin Cultured Vagal Afferent Neurons as Sensors for Intestinal Effector Molecules Journal Article biosensors, 2023. @article{Girardi2023, title = {Cultured Vagal Afferent Neurons as Sensors for Intestinal Effector Molecules}, author = {Gregory Girardi and Danielle Zumpano and Noah Goshi and Helen Raybould and Erkin Seker}, url = {https://www.mdpi.com/2079-6374/13/6/601}, doi = {10.3390/bios13060601}, year = {2023}, date = {2023-05-31}, journal = {biosensors}, abstract = {The gut–brain axis embodies the bi-directional communication between the gastrointestinal tract and the central nervous system (CNS), where vagal afferent neurons (VANs) serve as sensors for a variety of gut-derived signals. The gut is colonized by a large and diverse population of microorganisms that communicate via small (effector) molecules, which also act on the VAN terminals situated in the gut viscera and consequently influence many CNS processes. However, the convoluted in vivo environment makes it difficult to study the causative impact of the effector molecules on VAN activation or desensitization. Here, we report on a VAN culture and its proof-of-principle demonstration as a cell-based sensor to monitor the influence of gastrointestinal effector molecules on neuronal behavior. We initially compared the effect of surface coatings (poly-L-lysine vs. Matrigel) and culture media composition (serum vs. growth factor supplement) on neurite growth as a surrogate of VAN regeneration following tissue harvesting, where the Matrigel coating, but not the media composition, played a significant role in the increased neurite growth. We then used both live-cell calcium imaging and extracellular electrophysiological recordings to show that the VANs responded to classical effector molecules of endogenous and exogenous origin (cholecystokinin serotonin and capsaicin) in a complex fashion. We expect this study to enable platforms for screening various effector molecules and their influence on VAN activity, assessed by their information-rich electrophysiological fingerprints.}, keywords = {}, pubstate = {published}, tppubtype = {article} } The gut–brain axis embodies the bi-directional communication between the gastrointestinal tract and the central nervous system (CNS), where vagal afferent neurons (VANs) serve as sensors for a variety of gut-derived signals. The gut is colonized by a large and diverse population of microorganisms that communicate via small (effector) molecules, which also act on the VAN terminals situated in the gut viscera and consequently influence many CNS processes. However, the convoluted in vivo environment makes it difficult to study the causative impact of the effector molecules on VAN activation or desensitization. Here, we report on a VAN culture and its proof-of-principle demonstration as a cell-based sensor to monitor the influence of gastrointestinal effector molecules on neuronal behavior. We initially compared the effect of surface coatings (poly-L-lysine vs. Matrigel) and culture media composition (serum vs. growth factor supplement) on neurite growth as a surrogate of VAN regeneration following tissue harvesting, where the Matrigel coating, but not the media composition, played a significant role in the increased neurite growth. We then used both live-cell calcium imaging and extracellular electrophysiological recordings to show that the VANs responded to classical effector molecules of endogenous and exogenous origin (cholecystokinin serotonin and capsaicin) in a complex fashion. We expect this study to enable platforms for screening various effector molecules and their influence on VAN activity, assessed by their information-rich electrophysiological fingerprints. |
  | Xu, He Jax; Yao, Yao; Yao, Fenyong; Chen, Jiehui; Li, Meishi; Yang, Xianfa; Li, Sheng; Lu, Fangru; Hu, Ping; He, Shuijin; Peng, Guangdun; Jing, Naihe Generation of functional posterior spinal motor neurons from hPSCs-derived human spinal cord neural progenitor cells Journal Article Cell Regeneration, 2023. @article{Xu2023, title = {Generation of functional posterior spinal motor neurons from hPSCs-derived human spinal cord neural progenitor cells}, author = {He Jax Xu and Yao Yao and Fenyong Yao and Jiehui Chen and Meishi Li and Xianfa Yang and Sheng Li and Fangru Lu and Ping Hu and Shuijin He and Guangdun Peng and Naihe Jing}, url = {https://cellregeneration.springeropen.com/articles/10.1186/s13619-023-00159-6}, doi = {10.1186/s13619-023-00159-6}, year = {2023}, date = {2023-03-23}, journal = {Cell Regeneration}, abstract = {Spinal motor neurons deficiency results in a series of devastating disorders such as amyotrophic lateral sclerosis (ALS), spinal muscular atrophy (SMA) and spinal cord injury (SCI). These disorders are currently incurable, while human pluripotent stem cells (hPSCs)-derived spinal motor neurons are promising but suffered from inappropriate regional identity and functional immaturity for the study and treatment of posterior spinal cord related injuries. In this study, we have established human spinal cord neural progenitor cells (hSCNPCs) via hPSCs differentiated neuromesodermal progenitors (NMPs) and demonstrated the hSCNPCs can be continuously expanded up to 40 passages. hSCNPCs can be rapidly differentiated into posterior spinal motor neurons with high efficiency. The functional maturity has been examined in detail. Moreover, a co-culture scheme which is compatible for both neural and muscular differentiation is developed to mimic the neuromuscular junction (NMJ) formation in vitro. Together, these studies highlight the potential avenues for generating clinically relevant spinal motor neurons and modeling neuromuscular diseases through our defined hSCNPCs.}, keywords = {}, pubstate = {published}, tppubtype = {article} } Spinal motor neurons deficiency results in a series of devastating disorders such as amyotrophic lateral sclerosis (ALS), spinal muscular atrophy (SMA) and spinal cord injury (SCI). These disorders are currently incurable, while human pluripotent stem cells (hPSCs)-derived spinal motor neurons are promising but suffered from inappropriate regional identity and functional immaturity for the study and treatment of posterior spinal cord related injuries. In this study, we have established human spinal cord neural progenitor cells (hSCNPCs) via hPSCs differentiated neuromesodermal progenitors (NMPs) and demonstrated the hSCNPCs can be continuously expanded up to 40 passages. hSCNPCs can be rapidly differentiated into posterior spinal motor neurons with high efficiency. The functional maturity has been examined in detail. Moreover, a co-culture scheme which is compatible for both neural and muscular differentiation is developed to mimic the neuromuscular junction (NMJ) formation in vitro. Together, these studies highlight the potential avenues for generating clinically relevant spinal motor neurons and modeling neuromuscular diseases through our defined hSCNPCs. |
  | Qian, Junming; Guan, Xiaonan; Xie, Bing; Xu, Chuanyun; Niu, Jacqueline; Tang, Xin; Li, Charles H; Colecraft, Henry M; Jaenisch, Rudolf; Liu, Shawn X Multiplex epigenome editing of MECP2 to rescue Rett syndrome neurons Journal Article Science Translational Medicine, 2023. @article{Qian2023, title = {Multiplex epigenome editing of MECP2 to rescue Rett syndrome neurons}, author = {Junming Qian and Xiaonan Guan and Bing Xie and Chuanyun Xu and Jacqueline Niu and Xin Tang and Charles H. Li and Henry M. Colecraft and Rudolf Jaenisch and X. Shawn Liu}, url = {https://www.science.org/doi/10.1126/scitranslmed.add4666}, doi = {10.1126/scitranslmed.add4666}, year = {2023}, date = {2023-01-18}, journal = {Science Translational Medicine}, abstract = {Rettsyndrome(RTT)isanX-linkedneurodevelopmental disorder caused byloss-of-function heterozygous mutationsofmethyl CpG-binding protein2(MECP2) ontheXchromosome inyoungfemales. Reactivationofthe silent wild-type MECP2 allelefromtheinactiveXchromosome (Xi)represents apromising therapeutic opportunity forfemale patients withRTT.Here,weapplied amultiple xepigenome editing approachtoreactivate MECP2 fromXiinRTThuman embryonicstemcells(hESCs) andderivedneurons.Demethyla tionofthe MECP2 promoter bydCas9-T et1withtarget single-guide RNAreactivatedMECP2 fromXiinRTThESCs without detectable off-target effects atthetranscriptional level.Neuronsderivedfrommethyla tion-edited RTThESCs maintained MECP2 reactivationandreversedthesmaller somasizeandelectrophysiological abnormalities, twohallmarks ofRTT.InRTTneurons,insulationofthemethyla tion-edited MECP2 locusbydCpf1-CT CF (acatalytically deadCpf1fusedwithCCCTC-binding factor)withtarget CRISPR RNAenhanced MECP2 reactivationandrescued RTT-relatedneuronaldefects, providing aproof-of-concept studyforepigenome editing to treatRTTandpotentially otherdominant X-linkeddiseases.}, keywords = {}, pubstate = {published}, tppubtype = {article} } Rettsyndrome(RTT)isanX-linkedneurodevelopmental disorder caused byloss-of-function heterozygous mutationsofmethyl CpG-binding protein2(MECP2) ontheXchromosome inyoungfemales. Reactivationofthe silent wild-type MECP2 allelefromtheinactiveXchromosome (Xi)represents apromising therapeutic opportunity forfemale patients withRTT.Here,weapplied amultiple xepigenome editing approachtoreactivate MECP2 fromXiinRTThuman embryonicstemcells(hESCs) andderivedneurons.Demethyla tionofthe MECP2 promoter bydCas9-T et1withtarget single-guide RNAreactivatedMECP2 fromXiinRTThESCs without detectable off-target effects atthetranscriptional level.Neuronsderivedfrommethyla tion-edited RTThESCs maintained MECP2 reactivationandreversedthesmaller somasizeandelectrophysiological abnormalities, twohallmarks ofRTT.InRTTneurons,insulationofthemethyla tion-edited MECP2 locusbydCpf1-CT CF (acatalytically deadCpf1fusedwithCCCTC-binding factor)withtarget CRISPR RNAenhanced MECP2 reactivationandrescued RTT-relatedneuronaldefects, providing aproof-of-concept studyforepigenome editing to treatRTTandpotentially otherdominant X-linkeddiseases. |
  | McSweeney, Danny; Gabriel, Rafael; Jin, Kang; Pang, Zhiping P; Aronow, Bruce; and Pak, ChangHui CASK loss of function differentially regulates neuronal maturation and synaptic function in human induced cortical excitatory neurons Journal Article iScience, 2022. @article{McSweeney2022b, title = {CASK loss of function differentially regulates neuronal maturation and synaptic function in human induced cortical excitatory neurons}, author = {Danny McSweeney and Rafael Gabriel and Kang Jin and Zhiping P. Pang and Bruce Aronow and and ChangHui Pak}, url = {https://www.cell.com/iscience/fulltext/S2589-0042(22)01459-6?_returnURL=https%3A%2F%2Flinkinghub.elsevier.com%2Fretrieve%2Fpii%2FS2589004222014596%3Fshowall%3Dtrue}, doi = {https://doi.org/10.1016/j.isci.2022.105187}, year = {2022}, date = {2022-10-21}, journal = {iScience}, abstract = {Loss-of-function (LOF) mutations in CASK cause severe developmental pheno- types, including microcephaly with pontine and cerebellar hypoplasia, X-linked in- tellectual disability, and autism. Unraveling the pathological mechanisms of CASK-related disorders has been challenging owing to limited human cellular models to study the dynamic roles of this molecule during neuronal maturation and synapse development. Here, we investigate cell-autonomous functions of CASK in cortical excitatory induced neurons (iNs) generated from CASK knockout (KO) isogenic human embryonic stem cells (hESCs) using gene expression, mor- phometrics, and electrophysiology. While immature CASK KO iNs show robust neuronal outgrowth, mature CASK KO iNs display severe defects in syn- aptic transmission and synchronized network activity without compromising neuronal morphology and synapse numbers. In the developing human cortical excitatory neurons, CASK functions to promote both structural integrity and establishment of cortical excitatory neuronal networks. These results lay the foundation for future studies identifying suppressors of such phenotypes rele- vant to human patients.}, keywords = {}, pubstate = {published}, tppubtype = {article} } Loss-of-function (LOF) mutations in CASK cause severe developmental pheno- types, including microcephaly with pontine and cerebellar hypoplasia, X-linked in- tellectual disability, and autism. Unraveling the pathological mechanisms of CASK-related disorders has been challenging owing to limited human cellular models to study the dynamic roles of this molecule during neuronal maturation and synapse development. Here, we investigate cell-autonomous functions of CASK in cortical excitatory induced neurons (iNs) generated from CASK knockout (KO) isogenic human embryonic stem cells (hESCs) using gene expression, mor- phometrics, and electrophysiology. While immature CASK KO iNs show robust neuronal outgrowth, mature CASK KO iNs display severe defects in syn- aptic transmission and synchronized network activity without compromising neuronal morphology and synapse numbers. In the developing human cortical excitatory neurons, CASK functions to promote both structural integrity and establishment of cortical excitatory neuronal networks. These results lay the foundation for future studies identifying suppressors of such phenotypes rele- vant to human patients. |
  | Habibey, Rouhollah; Striebel, Johannes; Schmieder, Felix; Czarske, Jürgen; Busskamp, Volker Long-term morphological and functional dynamics of human stem cell-derived neuronal networks on high-density micro-electrode arrays Journal Article Frontiers in Neuroscience, 2022. @article{Habibey2022, title = {Long-term morphological and functional dynamics of human stem cell-derived neuronal networks on high-density micro-electrode arrays}, author = {Rouhollah Habibey and Johannes Striebel and Felix Schmieder and Jürgen Czarske and Volker Busskamp}, url = {https://www.frontiersin.org/articles/10.3389/fnins.2022.951964/full}, doi = {10.3389/fnins.2022.951964}, year = {2022}, date = {2022-10-04}, journal = {Frontiers in Neuroscience}, abstract = {Comprehensive electrophysiological characterizations of human induced pluripotent stem cell (hiPSC)-derived neuronal networks are essential to determine to what extent these in vitro models recapitulate the functional features of in vivo neuronal circuits. High-density micro-electrode arrays (HD-MEAs) offer non-invasive recording with the best spatial and temporal resolution possible to date. For 3 months, we tracked the morphology and activity features of developing networks derived from a transgenic hiPSC line in which neurogenesis is inducible by neurogenic transcription factor overexpression. Our morphological data revealed large-scale structural changes from homogeneously distributed neurons in the first month to the formation of neuronal clusters over time. This led to a constant shift in position of neuronal cells and clusters on HD-MEAs and corresponding changes in spatial distribution of the network activity maps. Network activity appeared as scarce action potentials (APs), evolved as local bursts with longer duration and changed to network-wide synchronized bursts with higher frequencies but shorter duration over time, resembling the emerging burst features found in the developing human brain. Instantaneous firing rate data indicated that the fraction of fast spiking neurons (150–600 Hz) increases sharply after 63 days post induction (dpi). Inhibition of glutamatergic synapses erased burst features from network activity profiles and confirmed the presence of mature excitatory neurotransmission. The application of GABAergic receptor antagonists profoundly changed the bursting profile of the network at 120 dpi. This indicated a GABAergic switch from excitatory to inhibitory neurotransmission during circuit development and maturation. Our results suggested that an emerging GABAergic system at older culture ages is involved in regulating spontaneous network bursts. In conclusion, our data showed that long-term and continuous microscopy and electrophysiology readouts are crucial for a meaningful characterization of morphological and functional maturation in stem cell-derived human networks. Most importantly, assessing the level and duration of functional maturation is key to subject these human neuronal circuits on HD-MEAs for basic and biomedical applications.}, keywords = {}, pubstate = {published}, tppubtype = {article} } Comprehensive electrophysiological characterizations of human induced pluripotent stem cell (hiPSC)-derived neuronal networks are essential to determine to what extent these in vitro models recapitulate the functional features of in vivo neuronal circuits. High-density micro-electrode arrays (HD-MEAs) offer non-invasive recording with the best spatial and temporal resolution possible to date. For 3 months, we tracked the morphology and activity features of developing networks derived from a transgenic hiPSC line in which neurogenesis is inducible by neurogenic transcription factor overexpression. Our morphological data revealed large-scale structural changes from homogeneously distributed neurons in the first month to the formation of neuronal clusters over time. This led to a constant shift in position of neuronal cells and clusters on HD-MEAs and corresponding changes in spatial distribution of the network activity maps. Network activity appeared as scarce action potentials (APs), evolved as local bursts with longer duration and changed to network-wide synchronized bursts with higher frequencies but shorter duration over time, resembling the emerging burst features found in the developing human brain. Instantaneous firing rate data indicated that the fraction of fast spiking neurons (150–600 Hz) increases sharply after 63 days post induction (dpi). Inhibition of glutamatergic synapses erased burst features from network activity profiles and confirmed the presence of mature excitatory neurotransmission. The application of GABAergic receptor antagonists profoundly changed the bursting profile of the network at 120 dpi. This indicated a GABAergic switch from excitatory to inhibitory neurotransmission during circuit development and maturation. Our results suggested that an emerging GABAergic system at older culture ages is involved in regulating spontaneous network bursts. In conclusion, our data showed that long-term and continuous microscopy and electrophysiology readouts are crucial for a meaningful characterization of morphological and functional maturation in stem cell-derived human networks. Most importantly, assessing the level and duration of functional maturation is key to subject these human neuronal circuits on HD-MEAs for basic and biomedical applications. |
  | Buccino Alessio Paolo; Damart, Tanguy; Bartram Julian; Mandge Darshan; Xue Xiaohan; Zbili Mickael; Gänswein Tobias; Jaquier Aurélien; Emmenegger Vishalini; Markram Henry; Hierlemann Andreas; Van Geit Werner. A multi-modal fitting approach to construct single-neuron models with patch clamp and high-density microelectrode arrays Journal Article bioRxiv, 2022. @article{Buccino2022, title = {A multi-modal fitting approach to construct single-neuron models with patch clamp and high-density microelectrode arrays}, author = {Buccino, Alessio Paolo; Damart, Tanguy; Bartram, Julian; Mandge, Darshan; Xue, Xiaohan; Zbili, Mickael; Gänswein, Tobias; Jaquier, Aurélien; Emmenegger, Vishalini; Markram, Henry; Hierlemann, Andreas; Van Geit, Werner.}, doi = {https://doi.org/10.1101/2022.08.03.502468}, year = {2022}, date = {2022-08-11}, journal = {bioRxiv}, abstract = {In computational neuroscience, multicompartment models are among the most biophysically realistic representations of single neurons. Constructing such models usually involves the use of the patch-clamp technique to record somatic voltage signals under different experimental conditions. The experimental data are then used to fit the many parameters of the model. While patching of the soma is currently the gold-standard approach to build multicompartment models, several studies have also evidenced a richness of dynamics in dendritic and axonal sections. Recording from the soma alone makes it hard to observe and correctly parameterize the activity of non-somatic compartments. In order to provide a richer set of data as input to multicompartment models, we here investigate the combination of somatic patch-clamp recordings with recordings of high-density micro-electrode arrays (HD-MEAs). HD-MEAs enable the observation of extracellular potentials and neural activity of neuronal compartments at sub-cellular resolution. In this work, we introduce a novel framework to combine patch-clamp and HD-MEA data to construct multicompartment models. We first validate our method on a ground-truth model with known parameters and show that the use of features extracted from extracellular signals, in addition to intracellular ones, yields models enabling better fits than using intracellular features alone. We also demonstrate our procedure using experimental data by constructing cell models from in vitro cell cultures. The proposed multi-modal fitting procedure has the potential to augment the modeling efforts of the computational neuroscience community and to provide the field with neuronal models that are more realistic and can be better validated. Author Summary Multicompartment models are one of the most biophysically detailed representations of single neurons. The vast majority of these models are built using experimental data from somatic recordings. However, neurons are much more than just their soma and one needs recordings from distal neurites to build an accurate model. In this article, we combine the patch-clamp technique with extracellular high-density microelectrode arrays (HD-MEAs) to compensate this shortcoming. In fact, HD-MEAs readouts allow one to record the neuronal signal in the entire axonal arbor. We show that the proposed multi-modal strategy is superior to the use of patch clamp alone using an existing model as ground-truth. Finally, we show an application of this strategy on experimental data from cultured neurons.}, keywords = {}, pubstate = {published}, tppubtype = {article} } In computational neuroscience, multicompartment models are among the most biophysically realistic representations of single neurons. Constructing such models usually involves the use of the patch-clamp technique to record somatic voltage signals under different experimental conditions. The experimental data are then used to fit the many parameters of the model. While patching of the soma is currently the gold-standard approach to build multicompartment models, several studies have also evidenced a richness of dynamics in dendritic and axonal sections. Recording from the soma alone makes it hard to observe and correctly parameterize the activity of non-somatic compartments. In order to provide a richer set of data as input to multicompartment models, we here investigate the combination of somatic patch-clamp recordings with recordings of high-density micro-electrode arrays (HD-MEAs). HD-MEAs enable the observation of extracellular potentials and neural activity of neuronal compartments at sub-cellular resolution. In this work, we introduce a novel framework to combine patch-clamp and HD-MEA data to construct multicompartment models. We first validate our method on a ground-truth model with known parameters and show that the use of features extracted from extracellular signals, in addition to intracellular ones, yields models enabling better fits than using intracellular features alone. We also demonstrate our procedure using experimental data by constructing cell models from in vitro cell cultures. The proposed multi-modal fitting procedure has the potential to augment the modeling efforts of the computational neuroscience community and to provide the field with neuronal models that are more realistic and can be better validated. Author Summary Multicompartment models are one of the most biophysically detailed representations of single neurons. The vast majority of these models are built using experimental data from somatic recordings. However, neurons are much more than just their soma and one needs recordings from distal neurites to build an accurate model. In this article, we combine the patch-clamp technique with extracellular high-density microelectrode arrays (HD-MEAs) to compensate this shortcoming. In fact, HD-MEAs readouts allow one to record the neuronal signal in the entire axonal arbor. We show that the proposed multi-modal strategy is superior to the use of patch clamp alone using an existing model as ground-truth. Finally, we show an application of this strategy on experimental data from cultured neurons. |
  | Xue, Xiaohan; Buccino, Alessio Paolo; Kumar, Sreedhar Saseendran; Hierlemann, Andreas; Bartram, Julian Inferring monosynaptic connections from paired dendritic spine Ca2+ imaging and large-scale recording of extracellular spiking Journal Article Journal of Neural Engineering, 2022. @article{Xue2022b, title = {Inferring monosynaptic connections from paired dendritic spine Ca2+ imaging and large-scale recording of extracellular spiking}, author = {Xiaohan Xue and Alessio Paolo Buccino and Sreedhar Saseendran Kumar and Andreas Hierlemann and Julian Bartram}, doi = {https://doi.org/10.1088/1741-2552/ac8765}, year = {2022}, date = {2022-08-11}, journal = {Journal of Neural Engineering}, abstract = {Techniques to identify monosynaptic connections between neurons have been vital for neuroscience research, facilitating important advancements concerning network topology, synaptic plasticity, and synaptic integration, among others. Here, we introduce a novel approach to identify and monitor monosynaptic connections using high-resolution dendritic spine Ca2+ imaging combined with simultaneous large-scale recording of extracellular electrical activity by means of high-density microelectrode arrays (HD-MEAs). We introduce an easily adoptable analysis pipeline that associates the imaged spine with its presynaptic unit and test it on in vitro recordings. The method is further validated and optimized by simulating synaptically-evoked spine Ca2+ transients based on measured spike trains in order to obtain simulated ground-truth connections. The proposed approach offers unique advantages as i) it can be used to identify monosynaptic connections with an accurate localization of the synapse within the dendritic tree, ii) it provides precise information of presynaptic spiking, and iii) postsynaptic spine Ca2+ signals and, finally, iv) the non-invasive nature of the proposed method allows for long-term measurements. The analysis toolkit together with the rich data sets that were acquired are made publicly available for further exploration by the research community.}, keywords = {}, pubstate = {published}, tppubtype = {article} } Techniques to identify monosynaptic connections between neurons have been vital for neuroscience research, facilitating important advancements concerning network topology, synaptic plasticity, and synaptic integration, among others. Here, we introduce a novel approach to identify and monitor monosynaptic connections using high-resolution dendritic spine Ca2+ imaging combined with simultaneous large-scale recording of extracellular electrical activity by means of high-density microelectrode arrays (HD-MEAs). We introduce an easily adoptable analysis pipeline that associates the imaged spine with its presynaptic unit and test it on in vitro recordings. The method is further validated and optimized by simulating synaptically-evoked spine Ca2+ transients based on measured spike trains in order to obtain simulated ground-truth connections. The proposed approach offers unique advantages as i) it can be used to identify monosynaptic connections with an accurate localization of the synapse within the dendritic tree, ii) it provides precise information of presynaptic spiking, and iii) postsynaptic spine Ca2+ signals and, finally, iv) the non-invasive nature of the proposed method allows for long-term measurements. The analysis toolkit together with the rich data sets that were acquired are made publicly available for further exploration by the research community. |
  | Xu, Jax H; Yao, Yao; Yao, Fenyong; Chen, Jiehui; Li, Meishi; Xianfa, ; Yang, ; Li, Sheng; Lu, Fangru; Hu, Ping; He, Shuijin; Peng, Guangdun; Jing, Naihe Generation of functional posterior spinal motor neurons from hPSCs-derived human spinal cord neural progenitor cells Journal Article BioRxiv, 2022. @article{Xu2022, title = {Generation of functional posterior spinal motor neurons from hPSCs-derived human spinal cord neural progenitor cells}, author = {Jax H. Xu and Yao Yao and Fenyong Yao and Jiehui Chen and Meishi Li and Xianfa and Yang and Sheng Li and Fangru Lu and Ping Hu and Shuijin He and Guangdun Peng and Naihe Jing}, url = {https://www.biorxiv.org/content/10.1101/2022.06.26.495599v1}, doi = {https://doi.org/10.1101/2022.06.26.495599}, year = {2022}, date = {2022-06-27}, journal = {BioRxiv}, abstract = {Spinal motor neurons deficiency results in a series of devastating disorders such as amyotrophic lateral sclerosis (ALS), spinal muscular atrophy (SMA) and spinal cord injury (SCI). These disorders are currently incurable, while human pluripotent stem cells (hPSCs)-derived spinal motor neurons are promising but suffered from low-efficiency, functional immaturity and lacks of posterior cellular identity. In this study, we have established human spinal cord neural progenitor cells (hSCNPCs) via hPSCs differentiated neuromesodermal progenitors (NMPs) and demonstrated the hSCNPCs can be continuously expanded up to 40 passages. hSCNPCs can be rapidly differentiated into posterior spinal motor neurons with high efficiency. The functional maturity has been examined in detail. Moreover, a co-culture scheme which is compatible for both neural and muscular differentiation is developed to mimic the neuromuscular junction (NMJ) formation in vitro. Together, these studies highlight the potential avenues for generating clinically relevant spinal motor neurons and modeling neuromuscular diseases through our defined hSCNPCs.}, keywords = {}, pubstate = {published}, tppubtype = {article} } Spinal motor neurons deficiency results in a series of devastating disorders such as amyotrophic lateral sclerosis (ALS), spinal muscular atrophy (SMA) and spinal cord injury (SCI). These disorders are currently incurable, while human pluripotent stem cells (hPSCs)-derived spinal motor neurons are promising but suffered from low-efficiency, functional immaturity and lacks of posterior cellular identity. In this study, we have established human spinal cord neural progenitor cells (hSCNPCs) via hPSCs differentiated neuromesodermal progenitors (NMPs) and demonstrated the hSCNPCs can be continuously expanded up to 40 passages. hSCNPCs can be rapidly differentiated into posterior spinal motor neurons with high efficiency. The functional maturity has been examined in detail. Moreover, a co-culture scheme which is compatible for both neural and muscular differentiation is developed to mimic the neuromuscular junction (NMJ) formation in vitro. Together, these studies highlight the potential avenues for generating clinically relevant spinal motor neurons and modeling neuromuscular diseases through our defined hSCNPCs. |
  | Kubota, Tomoyuki; Takahashi, Hirokazu; and Nakajima, Kohei Unifying framework for information processing in stochastically driven dynamical systems Journal Article Physical Review Research, 2021. @article{Kubota2023, title = {Unifying framework for information processing in stochastically driven dynamical systems}, author = {Tomoyuki Kubota and Hirokazu Takahashi and and Kohei Nakajima}, url = {https://journals.aps.org/prresearch/abstract/10.1103/PhysRevResearch.3.043135}, doi = {https://doi.org/10.1103/PhysRevResearch.3.043135}, year = {2021}, date = {2021-11-23}, journal = {Physical Review Research}, abstract = {A dynamical system is an information processing apparatus that encodes input streams from the external environment to its state and processes them through state transitions. The information processing capacity (IPC) is an excellent tool that comprehensively evaluates these processed inputs, providing details of unknown information processing in black box systems; however, this measure can be applied only to time-invariant systems. This paper extends the applicable range to time-variant systems and further reveals that the IPC is equivalent to coefficients of polynomial chaos (PC) expansion in more general dynamical systems. To achieve this objective, we tackle three issues. First, we establish a connection between the IPC for time-invariant systems and PC expansion, which is a type of polynomial expansion using orthogonal functions of input history as bases. We prove that the IPC corresponds to the squared norm of the coefficient vector of the basis in the PC expansion. Second, we show that an input following an arbitrary distribution can be used for the IPC, removing previous restrictions to specific input distributions. Third, we extend the conventional orthogonal bases to functions of both time and input history and propose the IPC for time-variant systems. To show the significance of our approach, we demonstrate that our measure can reveal information representations in not only machine learning networks but also a real, cultured neural network. Our generalized measure paves the way for unveiling the information processing capabilities of a wide variety of physical dynamics which have been left behind in nature.}, keywords = {}, pubstate = {published}, tppubtype = {article} } A dynamical system is an information processing apparatus that encodes input streams from the external environment to its state and processes them through state transitions. The information processing capacity (IPC) is an excellent tool that comprehensively evaluates these processed inputs, providing details of unknown information processing in black box systems; however, this measure can be applied only to time-invariant systems. This paper extends the applicable range to time-variant systems and further reveals that the IPC is equivalent to coefficients of polynomial chaos (PC) expansion in more general dynamical systems. To achieve this objective, we tackle three issues. First, we establish a connection between the IPC for time-invariant systems and PC expansion, which is a type of polynomial expansion using orthogonal functions of input history as bases. We prove that the IPC corresponds to the squared norm of the coefficient vector of the basis in the PC expansion. Second, we show that an input following an arbitrary distribution can be used for the IPC, removing previous restrictions to specific input distributions. Third, we extend the conventional orthogonal bases to functions of both time and input history and propose the IPC for time-variant systems. To show the significance of our approach, we demonstrate that our measure can reveal information representations in not only machine learning networks but also a real, cultured neural network. Our generalized measure paves the way for unveiling the information processing capabilities of a wide variety of physical dynamics which have been left behind in nature. |
  | Greve, Frauke; Lichtenberg, Jan; Kirstein, Kay Uwe; Frey, Urs; Perriard, Jean Claude; Hierlemann, Andreas A perforated CMOS microchip platform for immobilization and activity monitoring of electrogenic cells Journal Article Journal of Micromechanics and Microengineering, 17 (3), pp. 462-471, 2007, ISSN: 0960-1317. @article{Greve2007, title = {A perforated CMOS microchip platform for immobilization and activity monitoring of electrogenic cells}, author = {Frauke Greve and Jan Lichtenberg and Kay Uwe Kirstein and Urs Frey and Jean Claude Perriard and Andreas Hierlemann}, url = {http://iopscience.iop.org/article/10.1088/0960-1317/17/3/007/}, doi = {10.1088/0960-1317/17/3/007}, issn = {0960-1317}, year = {2007}, date = {2007-01-30}, journal = {Journal of Micromechanics and Microengineering}, volume = {17}, number = {3}, pages = {462-471}, abstract = {CMOS-based microelectrode systems offer decisive advantages over conventional micro-electrode arrays, which include the possibility to perform on-chip signal conditioning or to efficiently use larger numbers of electrodes to obtain statistically relevant data, e.g., in pharmacological drug screening. A larger number of electrodes can only be realized with the help of on-chip multiplexing and readout schemes, which require integrated electronics. Another fundamental issue in performing high-fidelity recordings from electrogenic cells is a good electrical coupling between the cells and the microelectrodes, in particular, since the recorded extracellular signals are in the range of only 10–1000 µV. In this paper we present the first CMOS microelectrode system with integrated micromechanical cell-placement features fabricated in a commercial CMOS process with subsequent post-CMOS bulk micromachining. This new microdevice aims at enabling the precise placement of single cells in the center of the electrodes to ensure an efficient use of the available electrodes, even for low-density cell cultures. Small through-chip holes have been generated at the metal-electrode sites by using a combination of bulk micromachining and reactive-ion etching. These holes act as orifices so that cell immobilization can be achieved by means of pneumatic anchoring. The chip additionally hosts integrated circuitry, i.e., multiplexers to select the respective readout electrodes, an amplifier with selectable gain (2×, 10×, 100×), and a high-pass filter (100 Hz cut-off). In this paper we show that electrical signals from most of the electrodes can be recorded, even in low-density cultures of neonatal rat cardiomyocytes, by using perforated metal electrodes and by applying a small underpressure from the backside of the chip. The measurements evidenced that, in most cases, about 90% of the electrodes were covered with single cells, approximately 4% were covered with more than one cell due to clustering and approximately 6% were not covered with any cell, mostly as a consequence of orifice clogging. After 4 days of culturing, the cells were still in place on the electrodes so that the cell electrical activity could be measured using the on-chip circuitry. Measured signal amplitudes were in the range of 500–700 µV, while the input-referred noise of the readout was below 15 µVrms (100 Hz–4 kHz bandwidth). We report on the development and fabrication of this new cell-biological tool and present first results collected during the characterization and evaluation of the chip. The recordings of electrical potentials of neonatal rat cardiomyocytes after several days in vitro, which, on the one hand, were conventionally cultured (no pneumatic anchoring) and, on the other hand, were anchored and immobilized, will be detailed.}, keywords = {}, pubstate = {published}, tppubtype = {article} } CMOS-based microelectrode systems offer decisive advantages over conventional micro-electrode arrays, which include the possibility to perform on-chip signal conditioning or to efficiently use larger numbers of electrodes to obtain statistically relevant data, e.g., in pharmacological drug screening. A larger number of electrodes can only be realized with the help of on-chip multiplexing and readout schemes, which require integrated electronics. Another fundamental issue in performing high-fidelity recordings from electrogenic cells is a good electrical coupling between the cells and the microelectrodes, in particular, since the recorded extracellular signals are in the range of only 10–1000 µV. In this paper we present the first CMOS microelectrode system with integrated micromechanical cell-placement features fabricated in a commercial CMOS process with subsequent post-CMOS bulk micromachining. This new microdevice aims at enabling the precise placement of single cells in the center of the electrodes to ensure an efficient use of the available electrodes, even for low-density cell cultures. Small through-chip holes have been generated at the metal-electrode sites by using a combination of bulk micromachining and reactive-ion etching. These holes act as orifices so that cell immobilization can be achieved by means of pneumatic anchoring. The chip additionally hosts integrated circuitry, i.e., multiplexers to select the respective readout electrodes, an amplifier with selectable gain (2×, 10×, 100×), and a high-pass filter (100 Hz cut-off). In this paper we show that electrical signals from most of the electrodes can be recorded, even in low-density cultures of neonatal rat cardiomyocytes, by using perforated metal electrodes and by applying a small underpressure from the backside of the chip. The measurements evidenced that, in most cases, about 90% of the electrodes were covered with single cells, approximately 4% were covered with more than one cell due to clustering and approximately 6% were not covered with any cell, mostly as a consequence of orifice clogging. After 4 days of culturing, the cells were still in place on the electrodes so that the cell electrical activity could be measured using the on-chip circuitry. Measured signal amplitudes were in the range of 500–700 µV, while the input-referred noise of the readout was below 15 µVrms (100 Hz–4 kHz bandwidth). We report on the development and fabrication of this new cell-biological tool and present first results collected during the characterization and evaluation of the chip. The recordings of electrical potentials of neonatal rat cardiomyocytes after several days in vitro, which, on the one hand, were conventionally cultured (no pneumatic anchoring) and, on the other hand, were anchored and immobilized, will be detailed. |
  | Heer, Flavio; Hafizovic, Sadik; Franks, Wendy; Blau, Axel; Ziegler, Christiane; Hierlemann, Andreas CMOS microelectrode array for bidirectional interaction with neuronal networks Journal Article IEEE Journal of Solid-State Circuits, 41 (7), pp. 1620-1629, 2006, ISSN: 0018-9200. @article{Heer2006, title = {CMOS microelectrode array for bidirectional interaction with neuronal networks}, author = {Flavio Heer and Sadik Hafizovic and Wendy Franks and Axel Blau and Christiane Ziegler and Andreas Hierlemann}, url = {http://ieeexplore.ieee.org/document/1644873/}, doi = {10.1109/JSSC.2006.873677}, issn = {0018-9200}, year = {2006}, date = {2006-07-07}, journal = {IEEE Journal of Solid-State Circuits}, volume = {41}, number = {7}, pages = {1620-1629}, abstract = {A CMOS metal-electrode-based micro system for bidirectional communication (stimulation and recording) with neuronal cells in vitro is presented. The chip overcomes the interconnect challenge that limits today's bidirectional microelectrode arrays. The microsystem has been fabricated in an industrial CMOS technology with several post-CMOS processing steps to realize 128 biocompatible electrodes and to ensure chip stability in physiological saline. The system comprises all necessary control circuitry and on-chip A/D and D/A conversion. A modular design has been implemented, where individual stimulation- and signal-conditioning circuitry units are associated with each electrode. Stimulation signals with a resolution of 8 bits can be sent to any subset of electrodes at a rate of 60 kHz, while all electrodes of the chip are continuously sampled at a rate of 20 kHz. The circuitry at each electrode can be individually reset to its operating point in order to suppress artifacts evoked by the stimulation pulses. Biological measurements from cultured neuronal networks originating from dissociated cortical tissue of fertilized chicken eggs with amplitudes of up to 500 muVpp are presented.}, keywords = {}, pubstate = {published}, tppubtype = {article} } A CMOS metal-electrode-based micro system for bidirectional communication (stimulation and recording) with neuronal cells in vitro is presented. The chip overcomes the interconnect challenge that limits today's bidirectional microelectrode arrays. The microsystem has been fabricated in an industrial CMOS technology with several post-CMOS processing steps to realize 128 biocompatible electrodes and to ensure chip stability in physiological saline. The system comprises all necessary control circuitry and on-chip A/D and D/A conversion. A modular design has been implemented, where individual stimulation- and signal-conditioning circuitry units are associated with each electrode. Stimulation signals with a resolution of 8 bits can be sent to any subset of electrodes at a rate of 60 kHz, while all electrodes of the chip are continuously sampled at a rate of 20 kHz. The circuitry at each electrode can be individually reset to its operating point in order to suppress artifacts evoked by the stimulation pulses. Biological measurements from cultured neuronal networks originating from dissociated cortical tissue of fertilized chicken eggs with amplitudes of up to 500 muVpp are presented. |
Contact Us
We would love to chat with you! Book a one-to-one call with one of our scientist to discuss how MaxWell Biosystems’ high-content electrophysiology solutions can bring new key insights to your project or request a quote.
 English
English


